!pip install "teeplot==1.4.2" "watermark==2.5.1" 2>&1 >/dev/null
%load_ext watermarkEnvironment Setup
import itertools as it
import functools
import random
from typing import Dict, List, Tuple
try:
import cupy as cp
except ImportError:
import numpy as cp
from IPython.display import display_html
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
from teeplot import teeplot as tp
from tqdm.auto import tqdm%watermark -diwmuv -ivLast updated: 2025-12-24T02:08:48.737837+00:00
Python implementation: CPython
Python version : 3.12.12
IPython version : 7.34.0
Compiler : GCC 11.4.0
OS : Linux
Release : 6.6.105+
Machine : x86_64
Processor : x86_64
CPU cores : 12
Architecture: 64bit
IPython : 7.34.0
cupy : 13.6.0
matplotlib: 3.10.0
numpy : 2.0.2
pandas : 2.2.2
seaborn : 0.13.2
teeplot : 1.4.2
tqdm : 4.67.1
Watermark: 2.5.1
use_cupy = True # use cupy backend (GPU), otherwise use numpy (CPU)
xp = [np, cp][use_cupy]Simulation Implementation
# State Constants
S: int = 0
I: int = 1
R: int = 2
@functools.lru_cache(maxsize=None)
def simulate(
N_SITES: int=2,
POP_SIZE: int=1_000_000,
BASE_B: float=0.3,
CONTACT_RATE: float=0.5,
RECOVERY_RATE: float=0.1,
MUTATION_RATE: float=1e-4,
# WANING_RATE: float=0.003,
WANING_RATE: float=0.016,
IMMUNE_STRENGTH: float=0.7,
WANED_STRENGTH: float=0.05,
N_STEPS: int=1_000,
seed: int=1,
) -> pd.DataFrame:
random.seed(seed)
np.random.seed(seed)
xp.random.seed(seed)
if N_SITES > 8:
raise NotImplementedError(
"current data types support only up to 8 sites",
)
def initialize_pop() -> Tuple[xp.ndarray, xp.ndarray, xp.ndarray]:
"""Initialize population statuses, genomes, and immune history."""
# pathogen_genomes: Bit k=0 is allele 2k, bit k=1 is allele 2k+1
pathogen_genomes = xp.zeros(shape=POP_SIZE, dtype=xp.uint8)
# host_immunities: Tracks 4 levels (3, 2, 1, 0) for each of the 2*N_SITES alleles
host_immunities = xp.full(
shape=(POP_SIZE, 2 * N_SITES), fill_value=3, dtype=xp.int8
)
# host_statuses: Current state (S, I, R)
host_statuses = xp.full(shape=POP_SIZE, fill_value=S, dtype=xp.uint8)
return host_statuses, pathogen_genomes, host_immunities
def infect_initial(
host_statuses: xp.ndarray,
pathogen_genomes: xp.ndarray,
seed_count: int = 100,
) -> Tuple[xp.ndarray, xp.ndarray]:
"""Seed the initial infection wave with the starting strain."""
host_statuses[:seed_count] = I
pathogen_genomes[:seed_count] = 0
return host_statuses, pathogen_genomes
def get_kappa(host_immunities: xp.ndarray) -> xp.ndarray:
"""Numerical value assigned to immunity levels (kappa)."""
kappas = xp.zeros_like(host_immunities, dtype=xp.float16)
# Level 2 or 1: Full immunity (kappa=1)
kappas[(host_immunities == 2) | (host_immunities == 1)] = 1.0
# Level 0: Waned immunity (kappa=nui)
kappas[host_immunities == 0] = WANED_STRENGTH
return kappas
def update_waning(host_immunities: xp.ndarray) -> xp.ndarray:
"""Transition immunity levels over time (2 -> 1 -> 0)."""
for level in [2, 1]:
mask = (host_immunities == level) & (xp.random.rand(*host_immunities.shape) < WANING_RATE)
host_immunities[mask] -= 1
return host_immunities
def update_recoveries(
host_statuses: xp.ndarray,
pathogen_genomes: xp.ndarray,
host_immunities: xp.ndarray,
) -> Tuple[xp.ndarray, xp.ndarray]:
"""Recover infected individuals and reset allele immunity to Level 2."""
inf_mask = host_statuses == I
rec_mask = inf_mask & (xp.random.rand(POP_SIZE) < RECOVERY_RATE)
indices = xp.where(rec_mask)[0]
if indices.size > 0:
g = pathogen_genomes[indices][:, None]
shifts = xp.arange(N_SITES, dtype=xp.uint64)
bits = (g >> shifts) & xp.uint8(1)
allele_indices = (2 * xp.arange(N_SITES) + bits).astype(int)
row_idx = xp.repeat(indices, N_SITES)
col_idx = allele_indices.flatten()
# Set to Level 2: "first stage of full immunity"
host_immunities[row_idx, col_idx] = 2
host_statuses[indices] = R
return host_statuses, host_immunities
def update_infections(
host_statuses: xp.ndarray,
pathogen_genomes: xp.ndarray,
host_immunities: xp.ndarray,
) -> Tuple[xp.ndarray, xp.ndarray, xp.ndarray]:
"""Vectorized transmission based on allele-specific susceptibility."""
infector_mask = host_statuses == I
num_infectors = int(xp.sum(infector_mask))
if num_infectors == 0:
return host_statuses, pathogen_genomes, host_immunities
targets = xp.random.randint(low=0, high=POP_SIZE, size=num_infectors, dtype=xp.uint32)
inf_genomes = pathogen_genomes[infector_mask]
# Map genomes to allele indices for susceptibility calculation
bits = (inf_genomes[:, None] >> xp.arange(N_SITES, dtype=xp.uint8)) & xp.uint8(1)
allele_indices = (2 * xp.arange(N_SITES) + bits).astype(int)
# Calculate susceptibility as product across all alleles
kappas = get_kappa(host_immunities[targets])
target_kappas = xp.take_along_axis(kappas, allele_indices, axis=1)
susc_factor = xp.prod(1.0 - (IMMUNE_STRENGTH * target_kappas), axis=1)
# Additive transmission factor
total_b = N_SITES * BASE_B
prob = total_b * CONTACT_RATE * susc_factor
success = (xp.random.rand(num_infectors) < prob) & (host_statuses[targets] != I)
new_inf_idx = targets[success]
if new_inf_idx.size > 0:
host_statuses[new_inf_idx] = I
new_genomes = inf_genomes[success]
# Mutation: flip one bit at a random site
mut_mask = xp.random.rand(new_inf_idx.size) < MUTATION_RATE
if xp.any(mut_mask):
num_mut = int(xp.sum(mut_mask))
flip_pos = xp.random.randint(low=0, high=N_SITES, size=num_mut).astype(xp.uint64)
new_genomes[mut_mask] ^= xp.uint64(1) << flip_pos
pathogen_genomes[new_inf_idx] = new_genomes
return host_statuses, pathogen_genomes, host_immunities
host_statuses, pathogen_genomes, host_immunities = initialize_pop()
host_statuses, pathogen_genomes = infect_initial(host_statuses, pathogen_genomes)
data_log: List[Dict[str, float]] = []
for t in tqdm(range(N_STEPS)):
host_statuses, host_immunities = update_recoveries(
host_statuses, pathogen_genomes, host_immunities
)
host_statuses, pathogen_genomes, host_immunities = update_infections(
host_statuses, pathogen_genomes, host_immunities
)
host_immunities = update_waning(host_immunities)
# 1. Strain Prevalence
inf_mask = host_statuses == I
counts_dict: Dict[str, float] = {}
if xp.any(inf_mask):
unique_g, counts = xp.unique(pathogen_genomes[inf_mask], return_counts=True)
# Fix: Site k bits -> Alleles 2k or 2k+1
bits = (unique_g[:, None] >> xp.arange(N_SITES, dtype=xp.uint8)) & xp.uint8(1)
alleles = 2 * xp.arange(N_SITES) + bits
strain_names = ["".join(map(str, row)) for row in alleles]
counts_dict = {f"Strain_{name}": c / POP_SIZE for name, c in zip(strain_names, counts)}
# 2. Host Immunity (Susceptibility per Allele)
# Calculate fraction of population susceptible to each allele j
pop_kappas = get_kappa(host_immunities)
# Susceptibility per allele = 1 - (mi * kappa)
pop_susc = xp.mean(1.0 - (IMMUNE_STRENGTH * pop_kappas), axis=0)
immunity_dict = {f"Susc_Allele_{j}": float(val) for j, val in enumerate(pop_susc)}
log_entry = {"Step": float(t), "Seed": seed}
log_entry.update(counts_dict)
log_entry.update(immunity_dict)
data_log.append(log_entry)
return pd.DataFrame(data_log).fillna(0)Plotting Implementation
def render_timeseries_plots(
df: pd.DataFrame,
suptitle: str,
teeplot_outattrs: dict,
) -> None:
for what, row in it.product(
["Susc", "Strain"],
["Seed", None],
):
data = df.filter(
regex=f"Step|Seed|{what}", axis=1
).melt(
id_vars=["Step", "Seed"], var_name="Class", value_name="Prevalence"
).astype(
{"Step": int, "Class": str, "Prevalence": float},
)
data["Ham. Wt."] = data["Class"].str.count("1|3|5|7|9")
palette = dict(zip(
data["Class"].unique(),
sns.color_palette("colorblind", len(data["Class"].unique())),
))
with tp.teed(
sns.relplot,
data=data,
x="Step",
y="Prevalence",
hue="Class",
col="Ham. Wt.",
row=row,
alpha=0.8,
dashes=False,
errorbar=("pi", 100),
err_kws=dict(alpha=0.1),
estimator=np.median,
facet_kws=dict(
margin_titles = True,
),
kind="line",
palette=palette,
teeplot_outattrs={
**teeplot_outattrs,
"what": what,
},
) as g:
g.map_dataframe(
sns.lineplot,
x="Step",
y="Prevalence",
hue="Class",
style="Seed",
alpha=0.7,
dashes=False,
errorbar=None,
legend=False,
linestyle=":",
linewidth=0.6,
palette=palette,
)
for ax in g.axes.flat:
ax.grid(True, alpha=0.3)
if what == "Strain":
g.set(yscale="log")
g.set(ylim=(1/POP_SIZE, 1.1))
g.figure.suptitle(suptitle)
if row is not None:
g.figure.subplots_adjust(hspace=0.16, top=0.9)
g.figure.set_size_inches(w=5, h=5)
else:
g.figure.subplots_adjust(top=0.7)
g.figure.set_size_inches(w=5, h=2)
sns.move_legend(g, "center left", bbox_to_anchor=(0.9, 0.5), frameon=False)Run Simulation and Render Plots across Condition Matrix
N_REP = 5
N_STEPS = 600
condition_matrix = it.product(
[5e-5, 1e-2], # MUTATION_RATE
[1_000_000, 10_000_000], # POP_SIZE
[2, 3], #N_SITES
)
for MUTATION_RATE, POP_SIZE, N_SITES in tqdm([*condition_matrix]):
suptitle = (
f"Pop Size: {POP_SIZE / 1_000_000}M., "
f"Mutation Rate: {MUTATION_RATE}, "
f"Num Sites: {N_SITES}"
)
display_html(f"<h2>{suptitle}</h2>", raw=True)
dfs = [
simulate(
MUTATION_RATE=MUTATION_RATE,
N_SITES=N_SITES,
N_STEPS=N_STEPS,
POP_SIZE=POP_SIZE,
seed=rep,
)
for rep in range(N_REP)
]
df = pd.concat(dfs)
render_timeseries_plots(
df=df,
suptitle=suptitle,
teeplot_outattrs={
"MUTATION_RATE".lower(): MUTATION_RATE,
"N_SITES".lower(): N_SITES,
"POP_SIZE".lower(): POP_SIZE,
},
)
Pop Size: 1.0M., Mutation Rate: 5e-05, Num Sites: 2
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=2+pop_size=1000000+row=seed+viz=relplot+what=Susc+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=2+pop_size=1000000+row=seed+viz=relplot+what=Susc+x=step+y=prevalence+ext=.png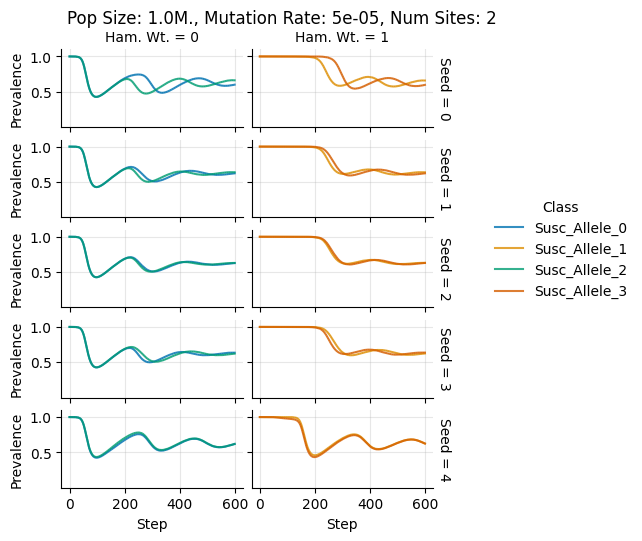
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=2+pop_size=1000000+viz=relplot+what=Susc+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=2+pop_size=1000000+viz=relplot+what=Susc+x=step+y=prevalence+ext=.png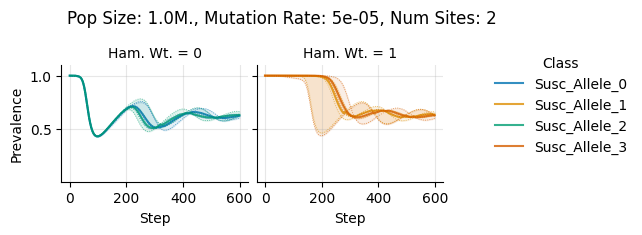
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=2+pop_size=1000000+row=seed+viz=relplot+what=Strain+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=2+pop_size=1000000+row=seed+viz=relplot+what=Strain+x=step+y=prevalence+ext=.png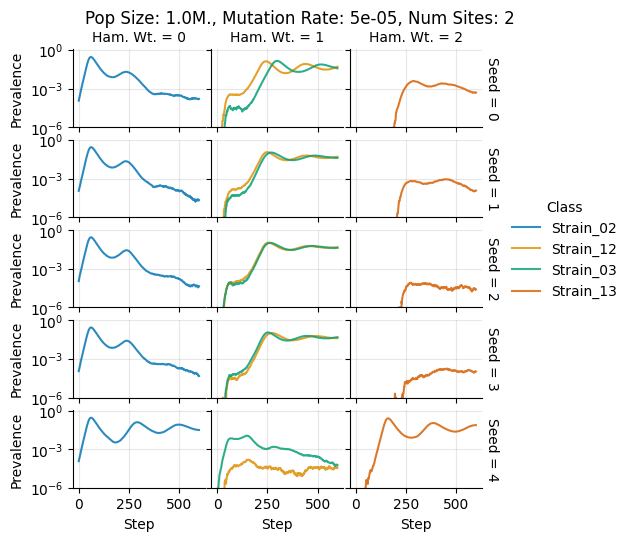
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=2+pop_size=1000000+viz=relplot+what=Strain+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=2+pop_size=1000000+viz=relplot+what=Strain+x=step+y=prevalence+ext=.png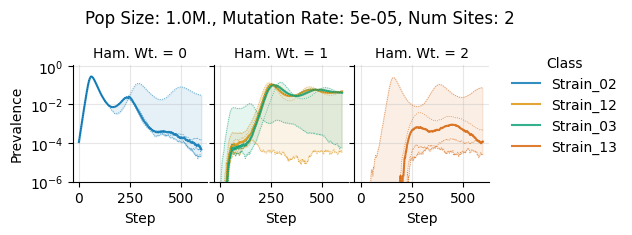
Pop Size: 1.0M., Mutation Rate: 5e-05, Num Sites: 3
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=3+pop_size=1000000+row=seed+viz=relplot+what=Susc+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=3+pop_size=1000000+row=seed+viz=relplot+what=Susc+x=step+y=prevalence+ext=.png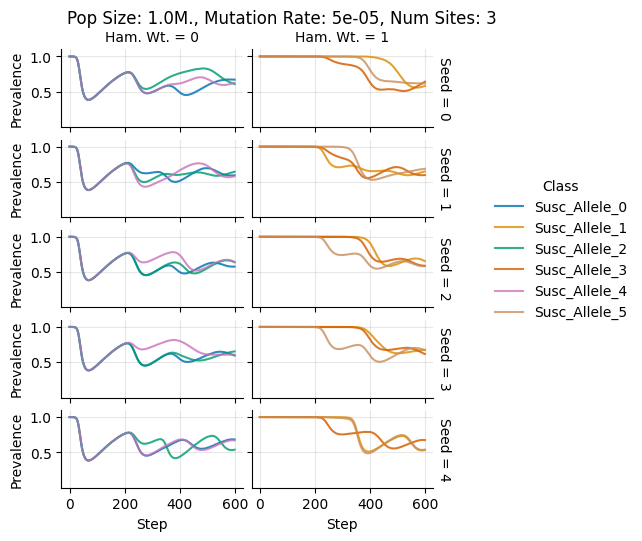
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=3+pop_size=1000000+viz=relplot+what=Susc+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=3+pop_size=1000000+viz=relplot+what=Susc+x=step+y=prevalence+ext=.png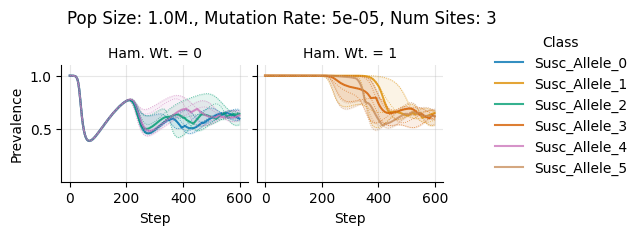
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=3+pop_size=1000000+row=seed+viz=relplot+what=Strain+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=3+pop_size=1000000+row=seed+viz=relplot+what=Strain+x=step+y=prevalence+ext=.png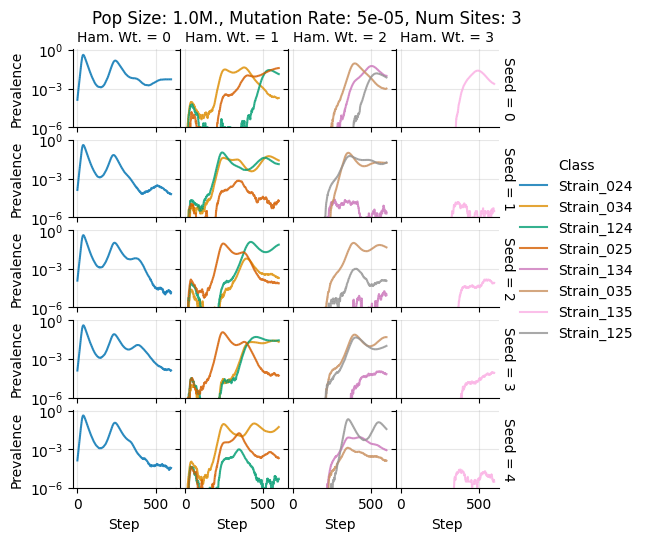
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=3+pop_size=1000000+viz=relplot+what=Strain+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=3+pop_size=1000000+viz=relplot+what=Strain+x=step+y=prevalence+ext=.png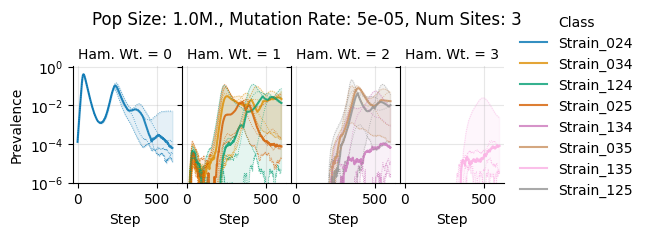
Pop Size: 10.0M., Mutation Rate: 5e-05, Num Sites: 2
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=2+pop_size=10000000+row=seed+viz=relplot+what=Susc+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=2+pop_size=10000000+row=seed+viz=relplot+what=Susc+x=step+y=prevalence+ext=.png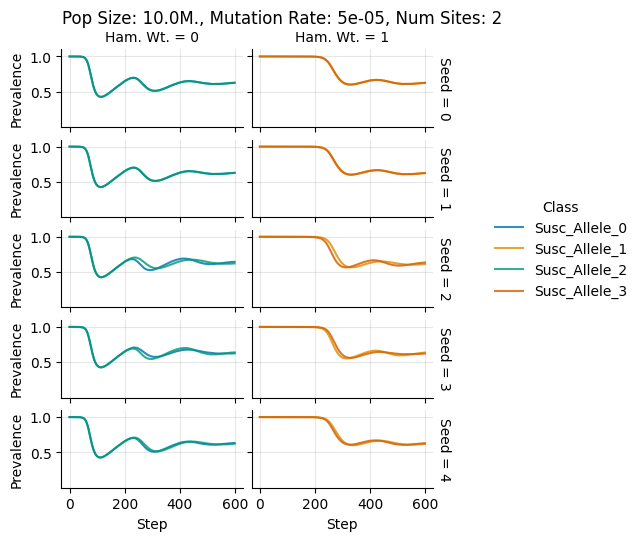
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=2+pop_size=10000000+viz=relplot+what=Susc+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=2+pop_size=10000000+viz=relplot+what=Susc+x=step+y=prevalence+ext=.png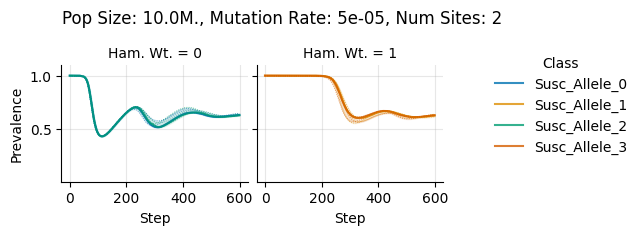
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=2+pop_size=10000000+row=seed+viz=relplot+what=Strain+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=2+pop_size=10000000+row=seed+viz=relplot+what=Strain+x=step+y=prevalence+ext=.png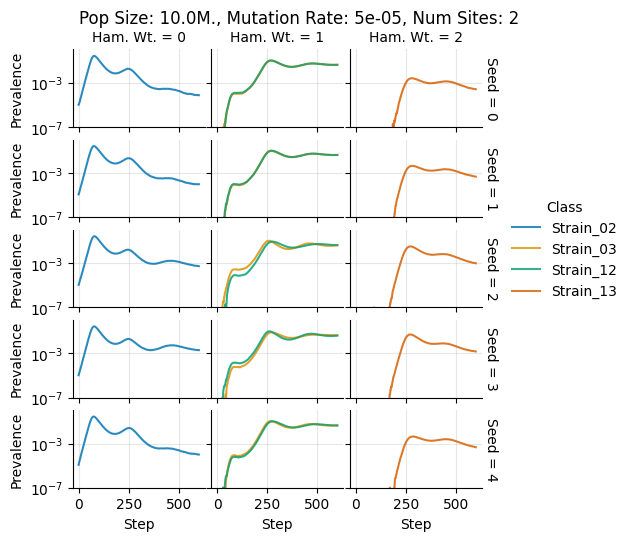
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=2+pop_size=10000000+viz=relplot+what=Strain+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=2+pop_size=10000000+viz=relplot+what=Strain+x=step+y=prevalence+ext=.png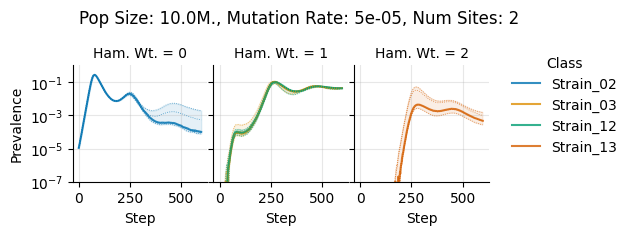
Pop Size: 10.0M., Mutation Rate: 5e-05, Num Sites: 3
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=3+pop_size=10000000+row=seed+viz=relplot+what=Susc+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=3+pop_size=10000000+row=seed+viz=relplot+what=Susc+x=step+y=prevalence+ext=.png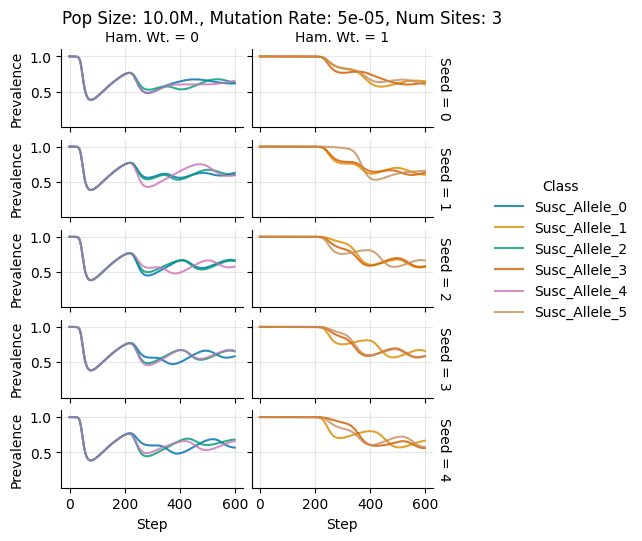
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=3+pop_size=10000000+viz=relplot+what=Susc+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=3+pop_size=10000000+viz=relplot+what=Susc+x=step+y=prevalence+ext=.png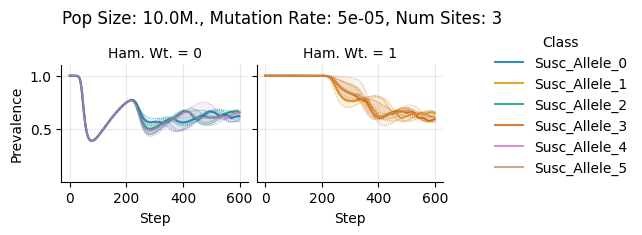
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=3+pop_size=10000000+row=seed+viz=relplot+what=Strain+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=3+pop_size=10000000+row=seed+viz=relplot+what=Strain+x=step+y=prevalence+ext=.png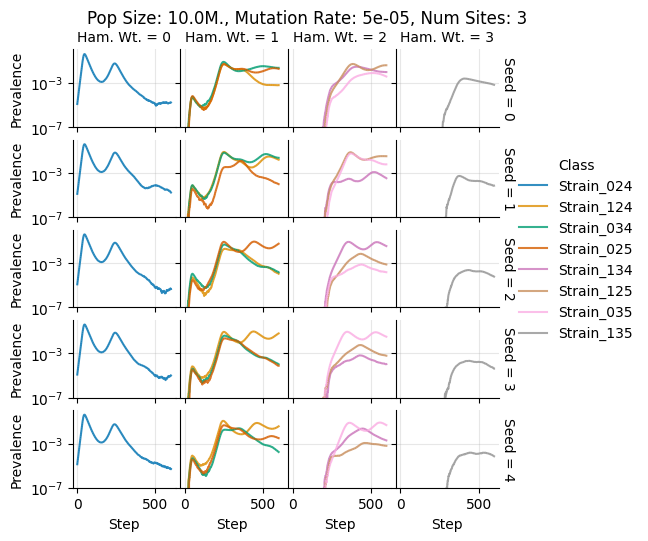
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=3+pop_size=10000000+viz=relplot+what=Strain+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=5e-05+n_sites=3+pop_size=10000000+viz=relplot+what=Strain+x=step+y=prevalence+ext=.png
Pop Size: 1.0M., Mutation Rate: 0.01, Num Sites: 2
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=2+pop_size=1000000+row=seed+viz=relplot+what=Susc+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=2+pop_size=1000000+row=seed+viz=relplot+what=Susc+x=step+y=prevalence+ext=.png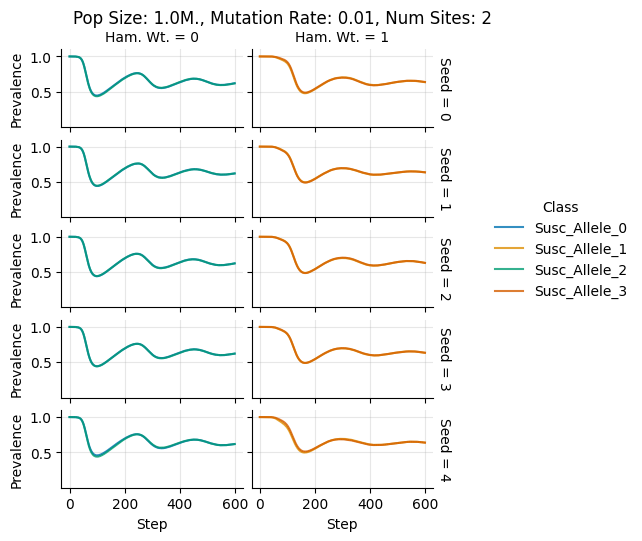
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=2+pop_size=1000000+viz=relplot+what=Susc+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=2+pop_size=1000000+viz=relplot+what=Susc+x=step+y=prevalence+ext=.png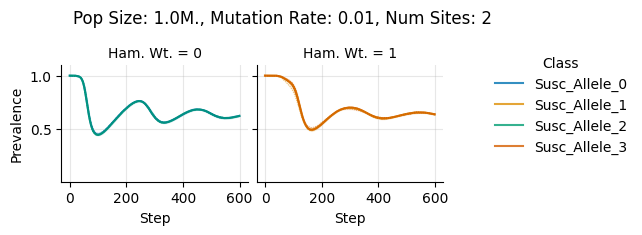
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=2+pop_size=1000000+row=seed+viz=relplot+what=Strain+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=2+pop_size=1000000+row=seed+viz=relplot+what=Strain+x=step+y=prevalence+ext=.png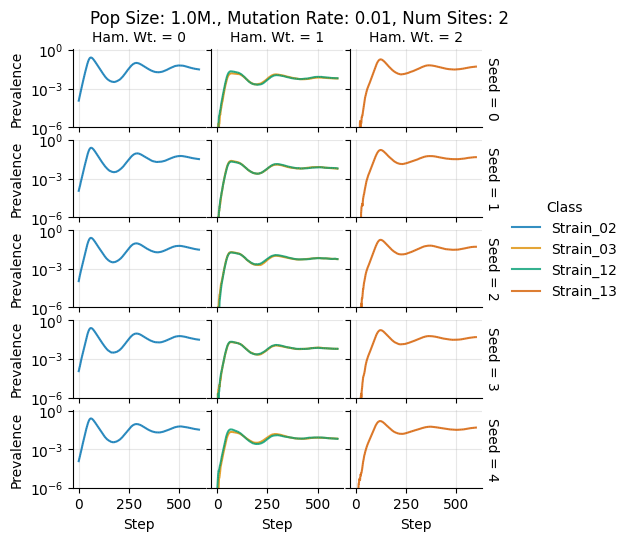
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=2+pop_size=1000000+viz=relplot+what=Strain+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=2+pop_size=1000000+viz=relplot+what=Strain+x=step+y=prevalence+ext=.png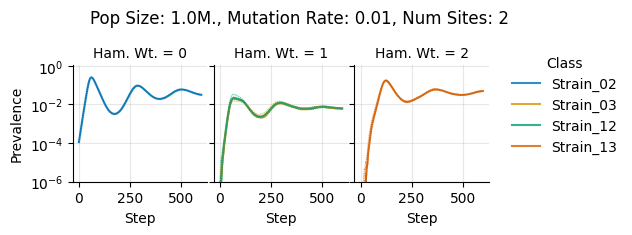
Pop Size: 1.0M., Mutation Rate: 0.01, Num Sites: 3
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=3+pop_size=1000000+row=seed+viz=relplot+what=Susc+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=3+pop_size=1000000+row=seed+viz=relplot+what=Susc+x=step+y=prevalence+ext=.png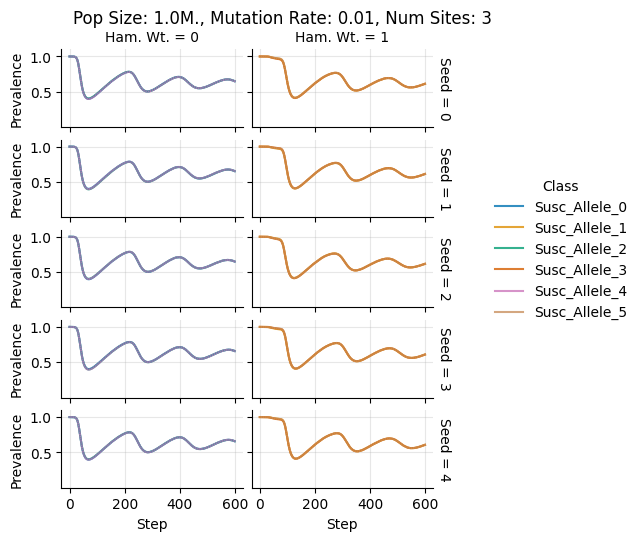
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=3+pop_size=1000000+viz=relplot+what=Susc+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=3+pop_size=1000000+viz=relplot+what=Susc+x=step+y=prevalence+ext=.png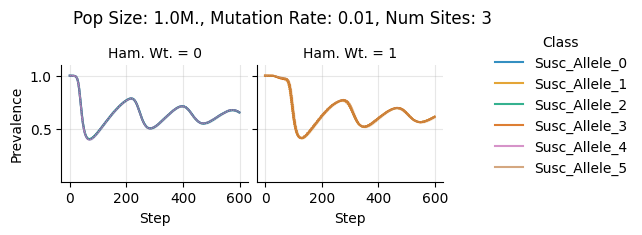
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=3+pop_size=1000000+row=seed+viz=relplot+what=Strain+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=3+pop_size=1000000+row=seed+viz=relplot+what=Strain+x=step+y=prevalence+ext=.png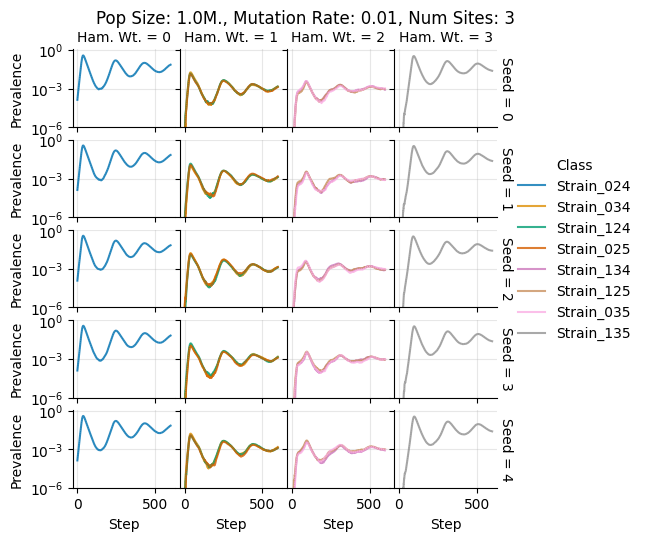
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=3+pop_size=1000000+viz=relplot+what=Strain+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=3+pop_size=1000000+viz=relplot+what=Strain+x=step+y=prevalence+ext=.png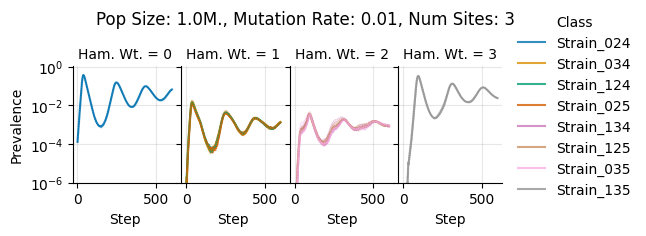
Pop Size: 10.0M., Mutation Rate: 0.01, Num Sites: 2
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=2+pop_size=10000000+row=seed+viz=relplot+what=Susc+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=2+pop_size=10000000+row=seed+viz=relplot+what=Susc+x=step+y=prevalence+ext=.png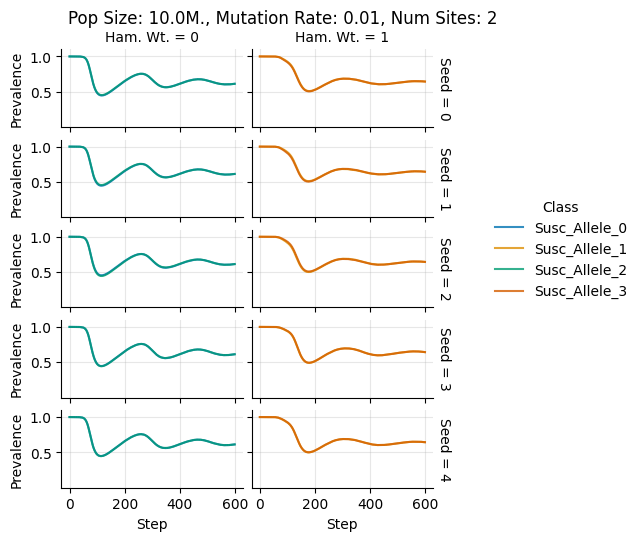
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=2+pop_size=10000000+viz=relplot+what=Susc+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=2+pop_size=10000000+viz=relplot+what=Susc+x=step+y=prevalence+ext=.png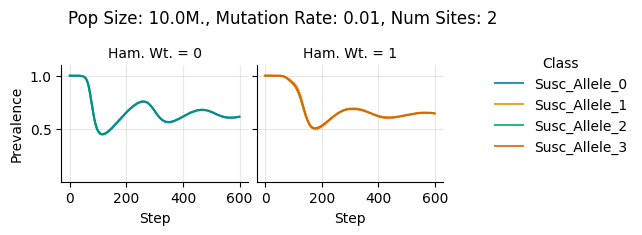
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=2+pop_size=10000000+row=seed+viz=relplot+what=Strain+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=2+pop_size=10000000+row=seed+viz=relplot+what=Strain+x=step+y=prevalence+ext=.png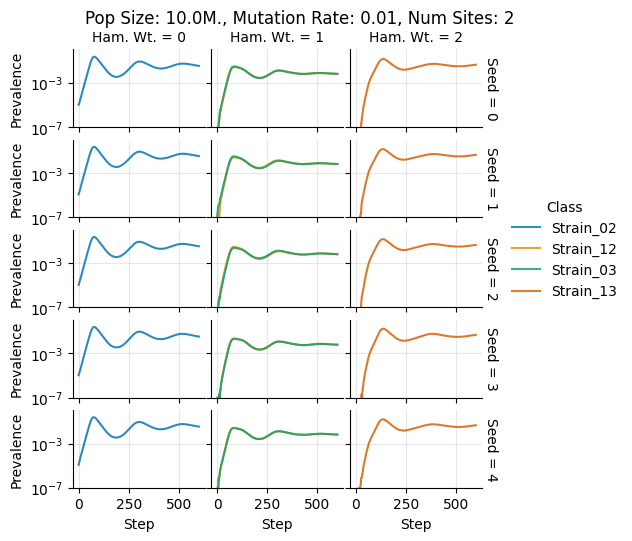
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=2+pop_size=10000000+viz=relplot+what=Strain+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=2+pop_size=10000000+viz=relplot+what=Strain+x=step+y=prevalence+ext=.png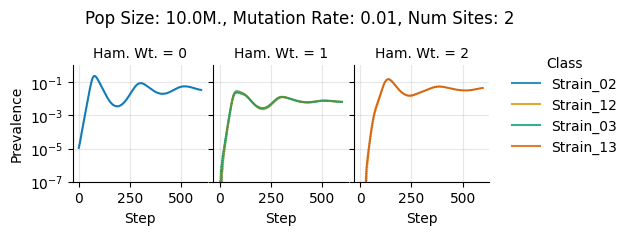
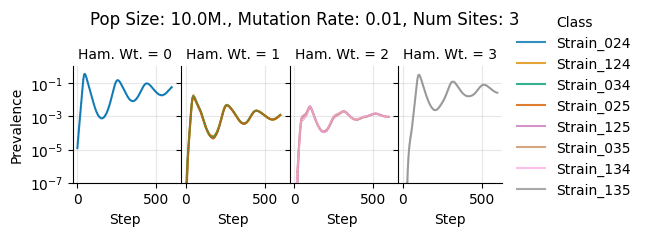
Pop Size: 10.0M., Mutation Rate: 0.01, Num Sites: 3
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=3+pop_size=10000000+row=seed+viz=relplot+what=Susc+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=3+pop_size=10000000+row=seed+viz=relplot+what=Susc+x=step+y=prevalence+ext=.png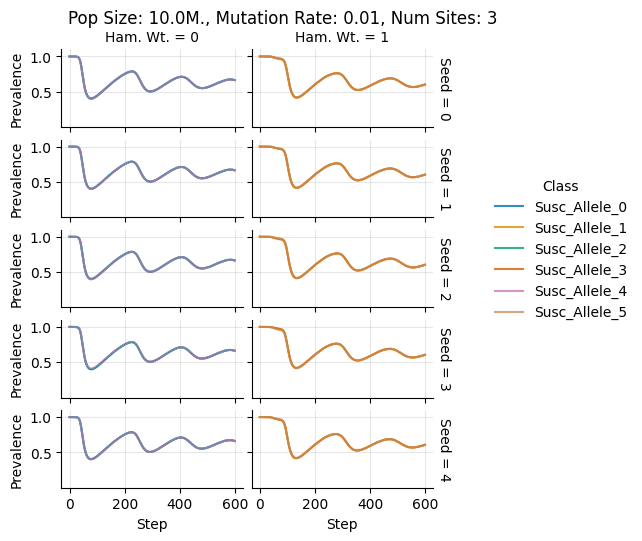
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=3+pop_size=10000000+viz=relplot+what=Susc+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=3+pop_size=10000000+viz=relplot+what=Susc+x=step+y=prevalence+ext=.png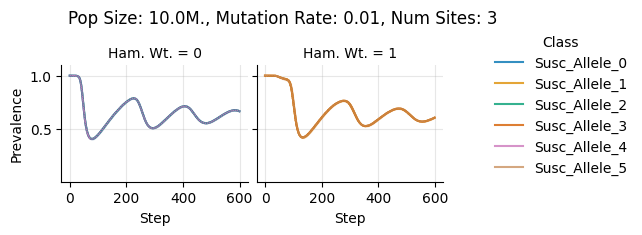
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=3+pop_size=10000000+row=seed+viz=relplot+what=Strain+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=3+pop_size=10000000+row=seed+viz=relplot+what=Strain+x=step+y=prevalence+ext=.png
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=3+pop_size=10000000+viz=relplot+what=Strain+x=step+y=prevalence+ext=.pdf
teeplots/col=ham-wt+hue=class+kind=line+mutation_rate=0.01+n_sites=3+pop_size=10000000+viz=relplot+what=Strain+x=step+y=prevalence+ext=.png